The SNP Detection System
The customer
The customer helps scientists and laboratories to conduct research and experiments in the field of life sciences. Their key services include next-generation sequencing, bioanalytical and mass spectrometry, as well as DNA sequencing. The customer turned to Altoros to develop a solution that would detect SNP in digitized DNA sequences saved in the FASTA/FASTQ format easier and less time-consuming.
The need
A common problem for researchers who work on genome analysis is the need to store and process terabytes of data fast. To address this issue, Altoros delivered an automated system for single-nucleotide polymorphism detection that provides better performance at a smaller cost. Deployed on Amazon public cloud, it was powered by Amazon Web Services and Amazon EMR. With this optimal solution our customer was able to process 150 GB of genome sequencing data within 24 hours and in the most cost-efficient manner.
Apart from building an algorithm for detecting SNP, we were to determine what hardware configuration could provide the required data processing speed.
The challenges
Apart from building an algorithm for detecting SNP, we were to determine what hardware configuration could provide the required data processing speed.
The solution
The team completed the following tasks for this project:
- Implementation of the data analysis algorithm. Our team designed a Web application to detect SNP and unite all tools required for genome analysis in one user-friendly interface. The software used Bowtie and SAMtools to align short DNA reads to the human genome and SOAPsnp to assemble consensus sequences and align raw sequencing reads on the known reference.
- Assessment of computation capacities. Our customer wanted to analyze heavy sets of sequencing data with an average size of 150 GB about 2-3 times a month. All computations had to be done within a maximum of 24 hours. We deployed the system on the Amazon cloud to keep the right balance between the cost of the solution and the throughput.
- Feasibility study and the system testing. Our team built a testing infrastructure using Amazon Web Services and Amazon Elastic MapReduce and provided a detailed report, where we indicated the cost of every solution depending on frequency of use, processing time, and amount of processed data.
- Building a private infrastructure. Although, the company was delighted with the results they achieved, they faced a new issue. The amount of data continued to grow and–eventually–they had to use AWS more frequently. It was decided to build a private infrastructure inside the customer’s laboratory.
The outcome
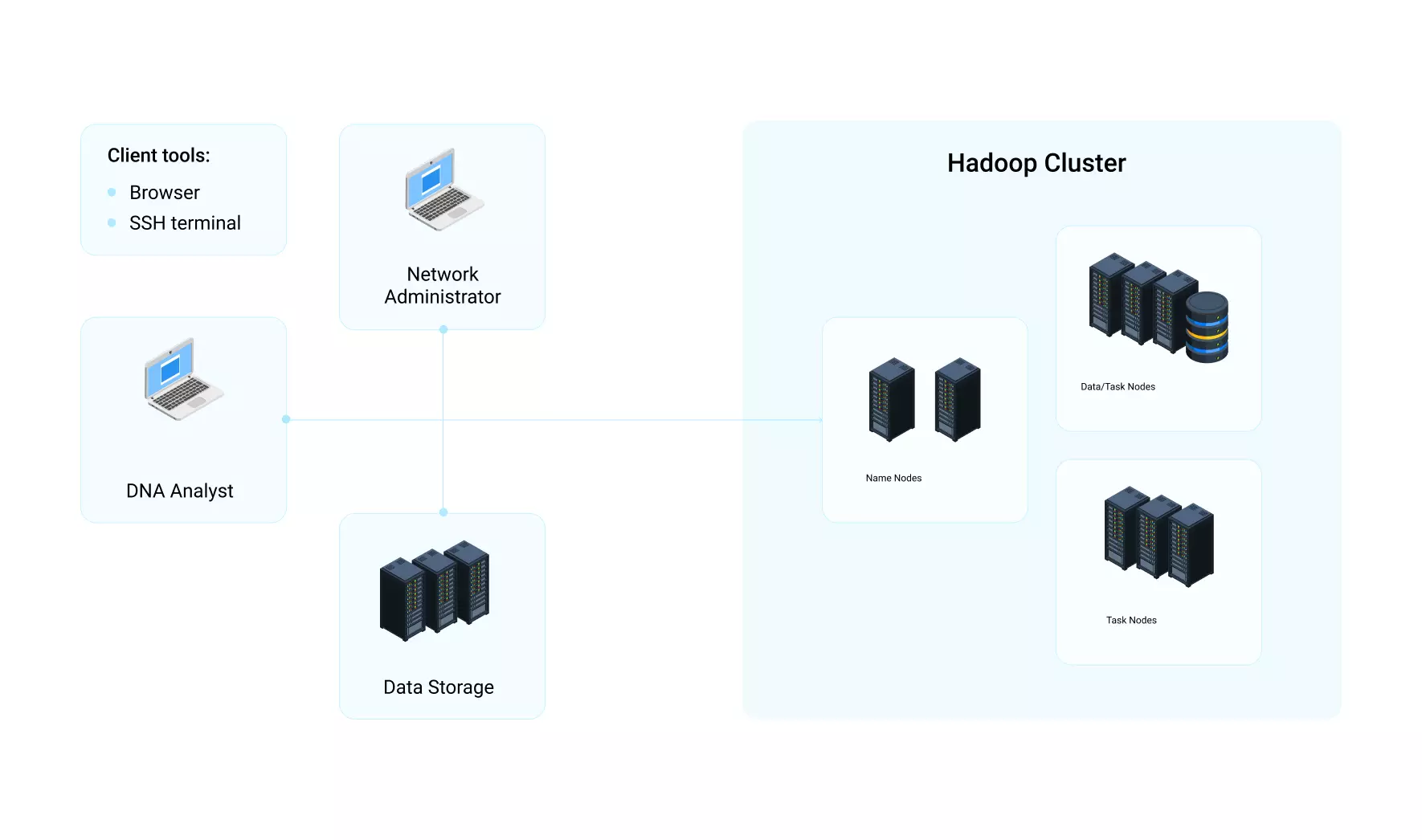
With the help of the automated SNP detection system, the biological laboratory of our customer managed to process 150GB of genome sequence data within 24 hours at minimum cost. We started with development of a prototype to test the possible deployment options and make sure the functionality works correctly. The system for SNP detection was later installed on the customer’s private distributed infrastructure and data processing was performed with Apache Hadoop.
Server Platform
Linux, Amazon Web Services
Client Platform/Application Server
Internet Explorer, Firefox, Safari, Chrome
Technologies
Map / Reduce, Java, HTML, Apache Hadoop, Amazon EMR
Programming languages
Perl, Java, Bash
Database, Storage
HDFS
Development Environment
Linux editors, Java IDE, Amazon AWS console
Seeking a solution like this?
Contact us and get a quote within 24 hours